Articles
- Page Path
- HOME > J Prev Med Public Health > Volume 45(4); 2012 > Article
-
Original Article
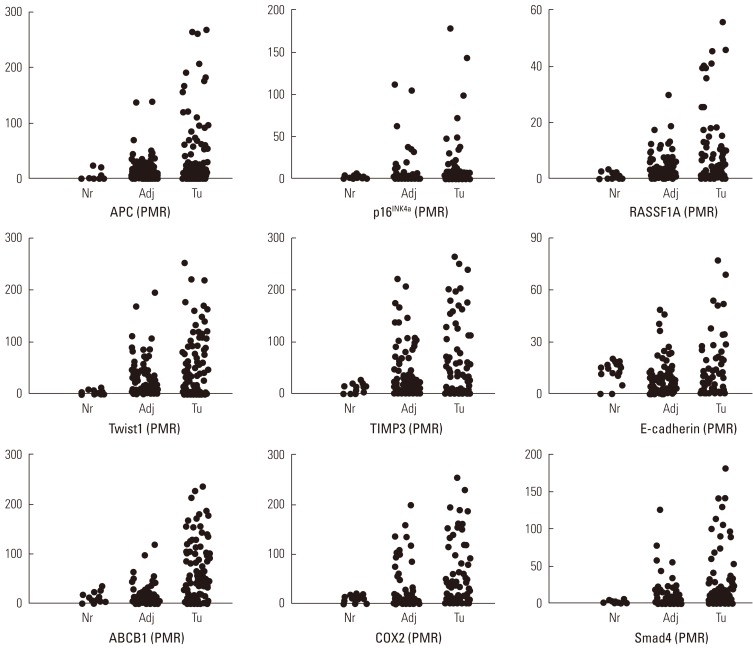
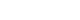
Quantitative Analysis of Cancer-associated Gene Methylation Connected to Risk Factors in Korean Colorectal Cancer Patients - Ho-Jin Kang1, Eun-Jeong Kim1, Byoung-Gwon Kim1, Chang-Hun You1, Sang-Yong Lee2, Dong-Il Kim3, Young-Seoub Hong1
-
Journal of Preventive Medicine and Public Health 2012;45(4):251-258.
DOI: https://doi.org/10.3961/jpmph.2012.45.4.251
Published online: July 31, 2012
- 9,270 Views
- 69 Download
- 11 Crossref
1Department of Preventive Medicine, Dong-A University College of Medicine, Busan, Korea.
2Institute of Forensic Medicine, Pusan National University School of Medicine, Busan, Korea.
3Department of Occupational Medicine, Kangbuk Samsung Hospital, Seoul, Korea.
- Corresponding author: Young-Seoub Hong, MD, PhD. 32 Daesingongwon-ro, Seo-gu, Busan 602-714, Korea. Tel: +82-51-240-2888, Fax: +82-51-253-5729, yshong@dau.ac.kr
• Received: September 26, 2011 • Accepted: March 14, 2012
Copyright © 2012 The Korean Society for Preventive Medicine
This is an Open Access article distributed under the terms of the Creative Commons Attribution Non-Commercial License (http://creativecommons.org/licenses/by-nc/3.0/) which permits unrestricted non-commercial use, distribution, and reproduction in any medium, provided the original work is properly cited.
Figure & Data
References
Citations
Citations to this article as recorded by 

- Relevance of gene mutations and methylation to the growth of pancreatic intraductal papillary mucinous neoplasms based on pyrosequencing
Go Asano, Katsuyuki Miyabe, Hiroyuki Kato, Michihiro Yoshida, Takeshi Sawada, Yasuyuki Okamoto, Hidenori Sahashi, Naoki Atsuta, Kenta Kachi, Akihisa Kato, Naruomi Jinno, Makoto Natsume, Yasuki Hori, Itaru Naitoh, Kazuki Hayashi, Yoichi Matsuo, Satoru Taka
Scientific Reports.2022;[Epub] CrossRef - Potential of RASSF1A promoter methylation as a biomarker for colorectal cancer: Meta-analysis and TCGA analysis
Fei Hu, Li Chen, Ming-Yu Bi, Ling Zheng, Ji-Xiang He, Ying-Ze Huang, Yu Zhang, Xue-Lian Zhang, Qiang Guo, Ying Luo, Wen-Ru Tang, Miao-Miao Sheng
Pathology - Research and Practice.2020; 216(8): 153009. CrossRef KLHL22 Regulates the EMT and Proliferation in Colorectal Cancer Cells in Part via the Wnt/β-Catenin Signaling Pathway
Yi Song, Huiping Yuan, Jia Wang, Yuhe Wu, Yuhong Xiao, Shengxun Mao
Cancer Management and Research.2020; Volume 12: 3981. CrossRef- RHBDF1 regulates APC-mediated stimulation of the epithelial-to-mesenchymal transition and proliferation of colorectal cancer cells in part via the Wnt/β-catenin signalling pathway
Huiping Yuan, Ran Wei, Yuhong Xiao, Yi Song, Jia Wang, Huihuan Yu, Ting Fang, Wei Xu, Shengxun Mao
Experimental Cell Research.2018; 368(1): 24. CrossRef - Smoking induces coordinated DNA methylation and gene expression changes in adipose tissue with consequences for metabolic health
Pei-Chien Tsai, Craig A. Glastonbury, Melissa N. Eliot, Sailalitha Bollepalli, Idil Yet, Juan E. Castillo-Fernandez, Elena Carnero-Montoro, Thomas Hardiman, Tiphaine C. Martin, Alice Vickers, Massimo Mangino, Kirsten Ward, Kirsi H. Pietiläinen, Panos Delo
Clinical Epigenetics.2018;[Epub] CrossRef - A novel discriminating colorectal cancer model for differentiating normal and tumor tissues
Xiaohui Sun, Yiping Tian, Qianqian Zheng, Ruizhi Zheng, Aifen Lin, Tianhui Chen, Yimin Zhu, Maode Lai
Epigenomics.2018; 10(11): 1463. CrossRef - APC hypermethylation for early diagnosis of colorectal cancer: a meta-analysis and literature review
Tie-Jun Liang, Hong-Xu Wang, Yan-Yan Zheng, Ying-Qing Cao, Xiaoyu Wu, Xin Zhou, Shu-Xiao Dong
Oncotarget.2017; 8(28): 46468. CrossRef - RETRACTED ARTICLE: Aberrant promoter methylation of RASSF1A gene may be correlated with colorectal carcinogenesis: a meta-analysis
He-Ling Wang, Yu Zhang, Peng Liu, Ping-Yi Zhou
Molecular Biology Reports.2014; 41(6): 3991. CrossRef - Role of CDH1 Promoter Methylation in Colorectal Carcinogenesis: A Meta-Analysis
Yu-Xi Li, Yao Lu, Chun-Yu Li, Peng Yuan, Shu-Sen Lin
DNA and Cell Biology.2014; 33(7): 455. CrossRef - Retracted: Promoter Methylation of theRASSF1AGene may Contribute to Colorectal Cancer Susceptibility: A Meta-Analysis of Cohort Studies
He-Ling Wang, Peng Liu, Ping-Yi Zhou, Yu Zhang
Annals of Human Genetics.2014; 78(3): 208. CrossRef - Hypermethylation ofTWIST1andNID2in Tumor Tissues and Voided Urine in Urinary Bladder Cancer Patients
Zeynep Yegin, Sezgin Gunes, Recep Buyukalpelli
DNA and Cell Biology.2013; 32(7): 386. CrossRef
 KSPM
KSPM

 PubReader
PubReader ePub Link
ePub Link Cite
Cite